The Botelho Lab @ Ryerson University in Toronto studies the molecular mechanisms that establish, maintain or adapt organelle identity and function. Organelles underpin everything that cells perform. Thus, understanding how organelles are made and function is essential to comprehend the underpinnings that drive life, health and disease- states. You are invited to explore and learn about our work and our energetic group. Please contact Dr. Roberto Botelho if interested in graduate or post-doctoral studies.
SELECT RESEARCH PROJECTS
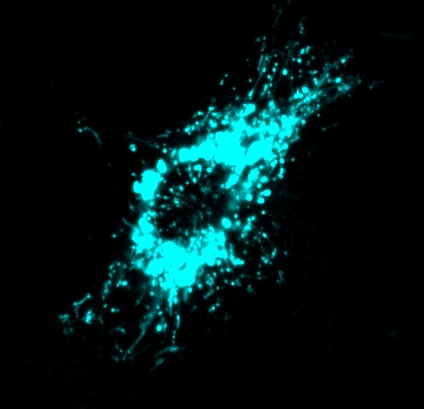
Lysosomes in dendritic cells exposed to monophosphoryl lipid A
mmune cells like macrophages and dendritic cells possess lysosomes that look like little globules under the fluorescence microscope. However, once these cells are activated by exposure to bacterial products or other immune triggers, these lysosomes change! For one thing, we showed that phagocytosis enhances expression of lysosomal proteins to bolster lysosomal activity (Gray et al.) . In other words, phagocytosis trains macrophages to become better killers. For another, lysosomes are transformed from many vesicular structures to a highly tubular network (see Saric et al. , for example). This transformation is also accompanied by expansion of the lysosome size through enhanced translational activity driven by mTORC1 (see Hipolito et al. ). Lysosome tubulation and expansion seems to play a role in antigen presentation, which the immune system then uses to “engineer” an attack plan on the source of antigens. For more information on this processes, see recent reviews from our lab in Cellular Microbiology (Hipolito et al LINK PMID: 29349904) and in Frontiers (Inpanathan and Botelho, LINK PMID: 31281815). Currently, we are how else lysosomes adapt during phagocyte activation, how lysosome remodelling matters for the immune response, and how the translational regulatory machinery aids to reorganize the endomembrane system.
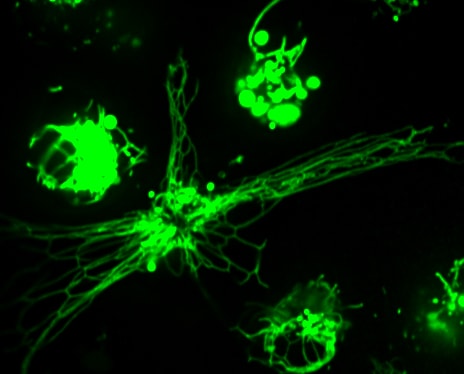
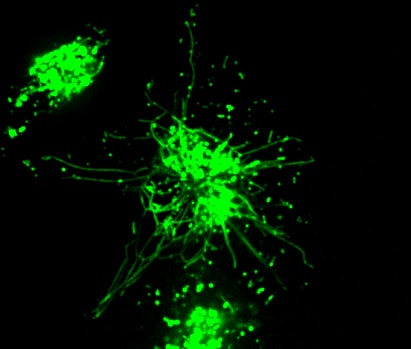
RAW 2FYVE GFP phagocytosing legionella filament
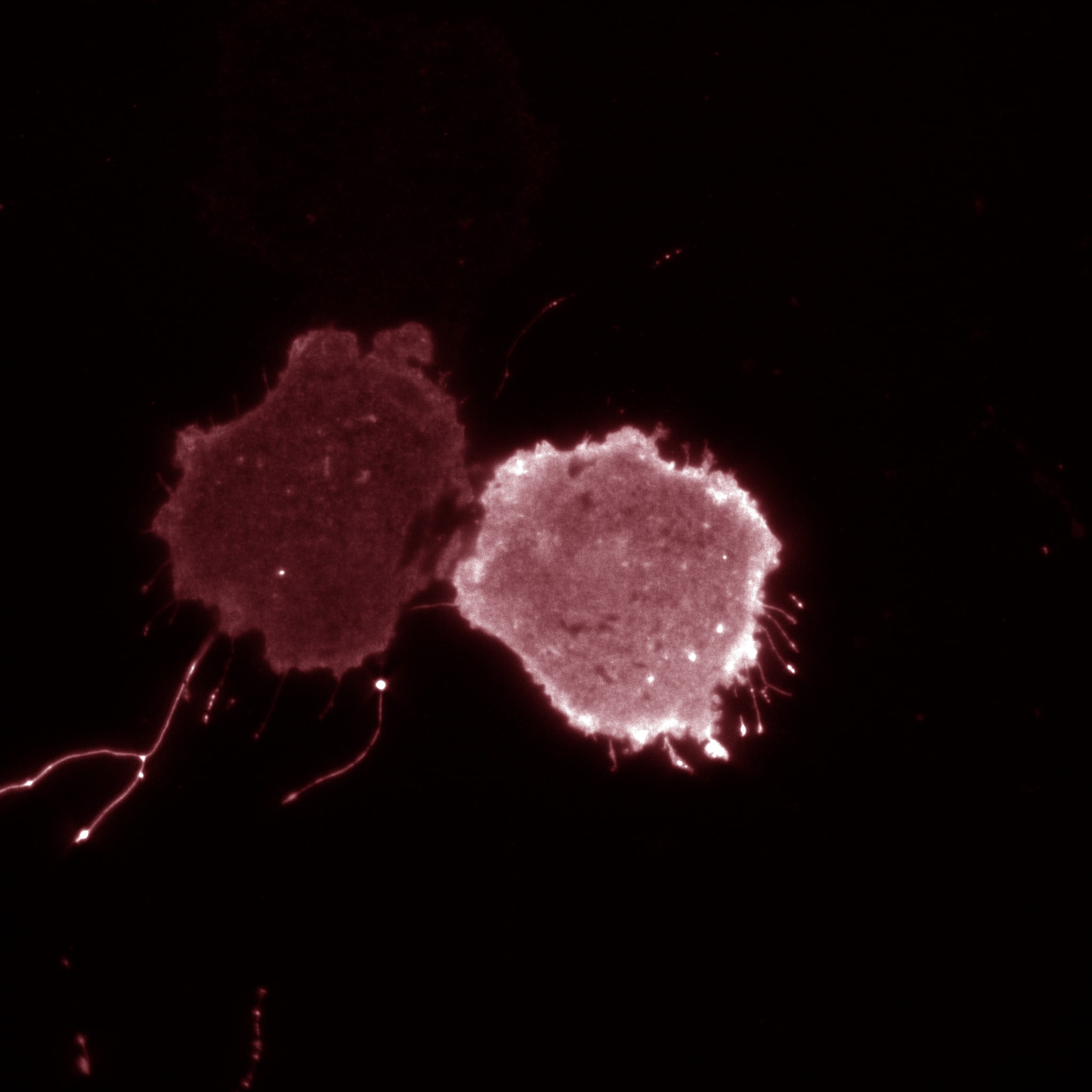
Macrophage eating filamentous Legionella (red). The macrophage is expressing 2FYVE-GFP, a fluorescent protein probe that tracks the dynamics of phosphatidylinositol-3- phosphate, a lipid that controls membrane trafficking. (In collaboration with Dr. Mauricio Terebiznik, U. of Toronto)
Macrophages, neutrophils and dendritic cells are immune cells that hunt down and engulf pathogens using their plasma membrane in a process called phagocytosis. After engulfment, the pathogen is sequestered in a new membrane-bound organelle called the phagosome. Importantly, the phagosome is then converted into a phagolysosome, a process known as phagosome maturation. This transformation changes the phagosome from an innocuous organelle to a very acidic and hydrolytic organelle that kills and digests the pathogen (see Gray & Botelho for recent review. Studying this system has two great advantages. First, this is a critically important process in the immune arsenal against pathogens – and one that can be hijacked by bugs like Salmonella and Mycobacterium. Second, the phagosome is a beautiful system to study organelle identity processes because we can track an organelle from birth (phagocytosis) to death (phagosome resolution) and study the changes in its identity over many hours. Currently, we are exploring how phosphoinositides regulate phagosome maturation (eg. Dayam et al., 2017), how cells adapt to phagocytosis (example: Gray et al. ), and how macrophage handles challenging particles like filamentous Legionella (eg. Naufer, Hipolito et al - a great collaboration with the Terebiznik lab at U of Toronto). Not yet published, we are also investigating what the heck happens to phagosomes after bacteria are digested!
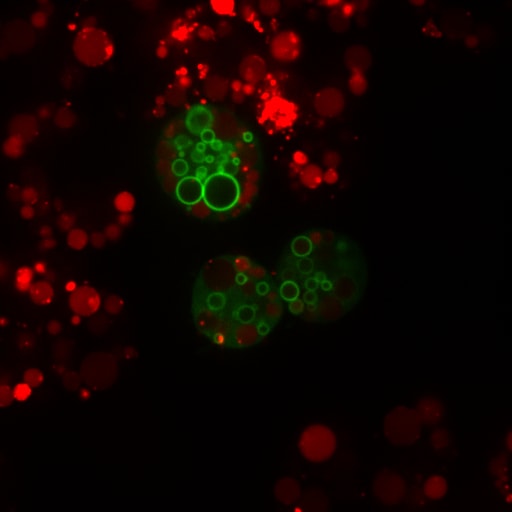
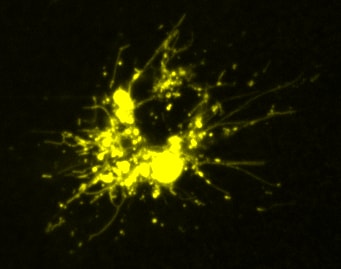
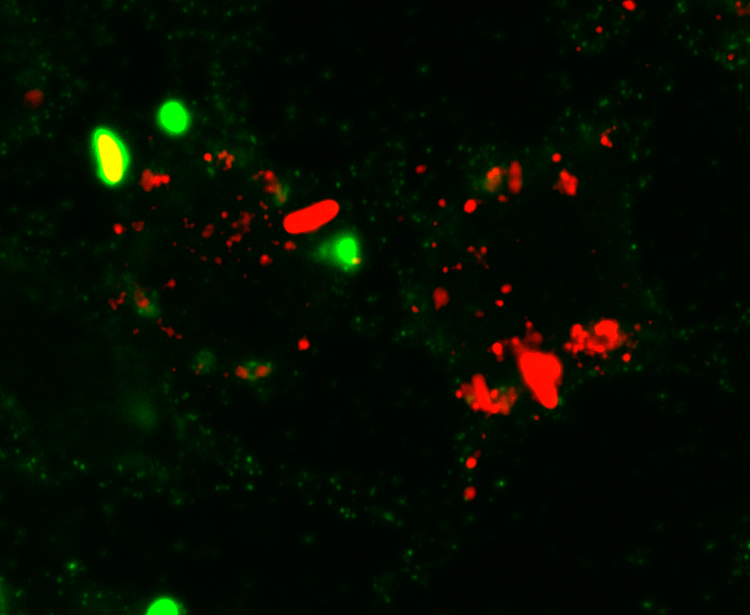
Cells treated with PIKfyve inhibitors display massively swollen lysosomes (red). Green is a new GFP-probe for phosphatidylinositol-3- phosphate.
We are investing resources to better understand the regulation and function of phosphatidylinositol-3,5-bisphosphate [PtdIns(3,5)P2], a phosphoinositide that controls properties of lysosomes and the yeast vacuole. As mentioned earlier, lysosomes are responsible for degrading macromolecules, organelles, and pathogens and recycle the raw materials to use as nutrients. They also can serve as storage organelles. When cells cannot synthesize PtdIns(3,5)P2, lysosomes become massively swollen, autophagic flux is impaired, phagosome maturation is arrested, even neutrophil chemotaxis is blunted, organisms fail to develop properly, there is a burst of inflammation, and neurological problems arise. So arguably, PtdIns(3,5)P2 kind of matters. For recent work on this lipid by our lab see Ho et al and Choy, Saffi et al. We are now trying to understand how lysosomes become massively enlarged and how this lipid coordinates a variety of seemingly unrelated processes to lysosomes including gene expression mechanisms.
We typically recognize that there are seven phosphoinositide (PIP) lipids based on the phosphorylation state of the headgroup (like PtdIns(3,5)P2 above). However, PIPs are phospholipids and thus contain two fatty acid tails. Interestingly, PIPs are enriched for two types of fatty acids: stearate, which is 18 carbons long and saturated, and arachidonic acid, an omega-6 lipid that is 20 carbons long and unsaturated. In collaboration with the Antonescu lab (link), we have been working to understand how cells establish this acyl profile and how this impacts cellular functions (see Bone et al., 2017 ). Current research projects are now focused on understanding i) how PIP acyl profile is important for cell signalling, cancer proliferation, and metastasis and ii) the mechanisms that establish PIP acyl profile. For more detailed discussion on these topics, please see Choy et al for a recent review on PIP biology.
SELECT PUBLICATION
For a full list of publications please refer to GOOGLE SCHOLAR or ORCID
Lysosome Fission: Planning for an Exit.
Saffi GT, Botelho RJ
Trends Cell Biol. 2019 Aug;29(8):635-646. doi: 10.1016/j.tcb.2019.05.003.
Abstract
Lysosomes are acidic and degradative organelles that receive and digest a plethora of molecular and particulate cargo delivered by endocytosis, autophagy, and phagocytosis. The mechanisms responsible for sorting, transporting, and ultimately delivering membranes and cargo to lysosomes through fusion have been intensely investigated. Much less is understood about lysosome fission, which is necessary to balance the incessant flow of cargo into lysosomes and maintain steady-state number, size, and function of lysosomes. Here, we review the emerging picture of how lipid signals, coat and adaptor proteins, and motor-cytoskeletal assemblies drive budding, tubulation, splitting, and 'kiss-and-run' events that enable fission and exit from lysosomes and related organelles.
The Lysosome Signaling Platform: Adapting With the Times
Inpanathan S, Botelho RJ
Front Cell Dev Biol. 2019 Jun 20;7:113. doi: 10.3389/fcell.2019.00113.
Lysosome enlargement during inhibition of the lipid kinase PIKfyve proceeds through lysosome coalescence
Choy CH, Saffi G, Gray MA, Wallace C, Dayam RM, Ou ZA, Lenk G, Puertollano R, Watkins SC, Botelho RJ
J Cell Biol. 2018 Jan 2;217(1):329-346. doi: 10.1083/jcb.201702179.
pH of endophagosomes controls association of their membranes with Vps34 and PtdIns(3)P level
Naufer A, Hipolito VEB, Ganesan S, Prashar A, Zaremberg V, Botelho RJ , Terebiznik MR
J Cell Biol. 2018 Jan 2;217(1):329-346. doi: 10.1083/jcb.201702179.
[1] Phosphoinositide Diversity, Distribution, and Effector Function: Stepping Out of the Box.
[2] Kinase PIKfyve Coordinates the Neutrophil Immune Response through the Activation of the Rac GTPase.
[3] The acyltransferase LYCAT controls specific phosphoinositides and related membrane traffic
[4] Phagocytosis Enhances Lysosomal and Bactericidal Properties by Activating the Transcription Factor TFEB.
[5] mTOR controls lysosome tubulation and antigen presentation in macrophages and dendritic cells.
[6] The Phosphoinositide-Gated Lysosomal Ca(2+) Channel, TRPML1, Is Required for Phagosome Maturation.
[7] The Fab1/PIKfyve phosphoinositide phosphate kinase is not necessary to maintain the pH of lysosomes and of the yeast vacuole.
[8] PIKfyve inhibition interferes with phagosome and endosome maturation in macrophages
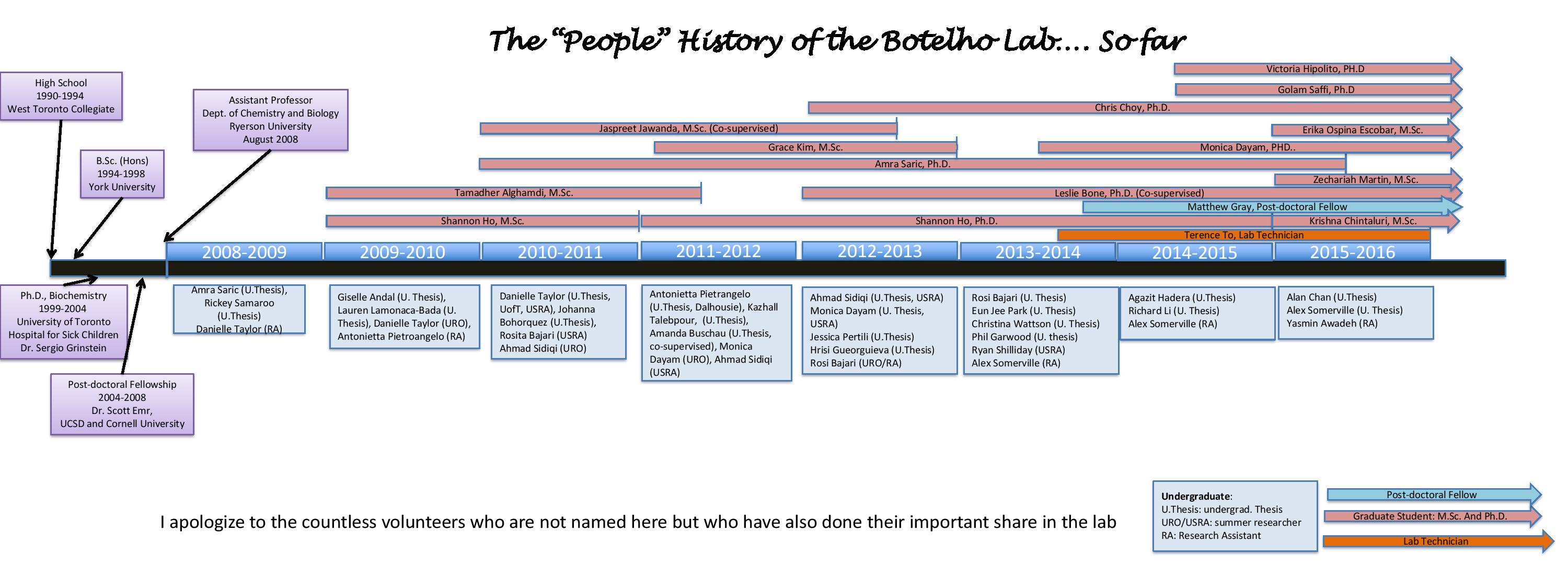
THE CREW
Crew

















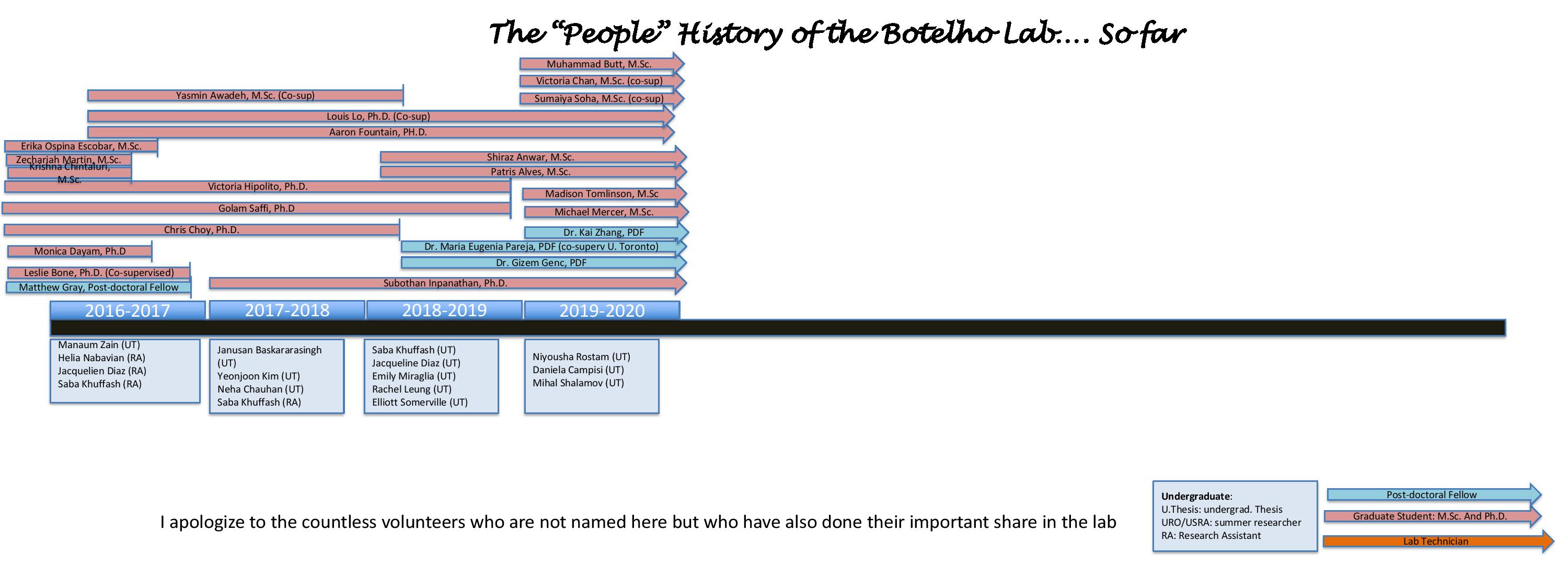
Alumni












Social Activities
From Halloween door competition to ski trips to ale-time and holidays parties. We try to do things together (though admittedly this requires a party planner!). Below shows some activities we have done over the years... Antonescu group is often a partner in crime.
Lab Environment

Earth, Canada, Toronto : The Botelho Lab is found in Downtown Toronto, also known as centre of the Universe (:p). That means you can travel the world within a few blocks, including eat burritos for breakfast, sashimi for lunch and vindaloo for dinner, go shopping, see a movie or a ballet, visit a museum, go skating, go for a walk to o chill by the lake or The Toronto Islands, take a boat cruise, climb the CN Tower, go see a game (even the Leafs), and feed pigeons and squirrels (if that's your thing).

Ryerson University and the Department of Chemistry and Biology are the academic and administrative home for the Botelho Lab. For quick facts on Ryerson and how much it has grown see here. Graduate students in the lab are enrolled in Molecular Science Graduate Program . I am always looking for committed and driven students

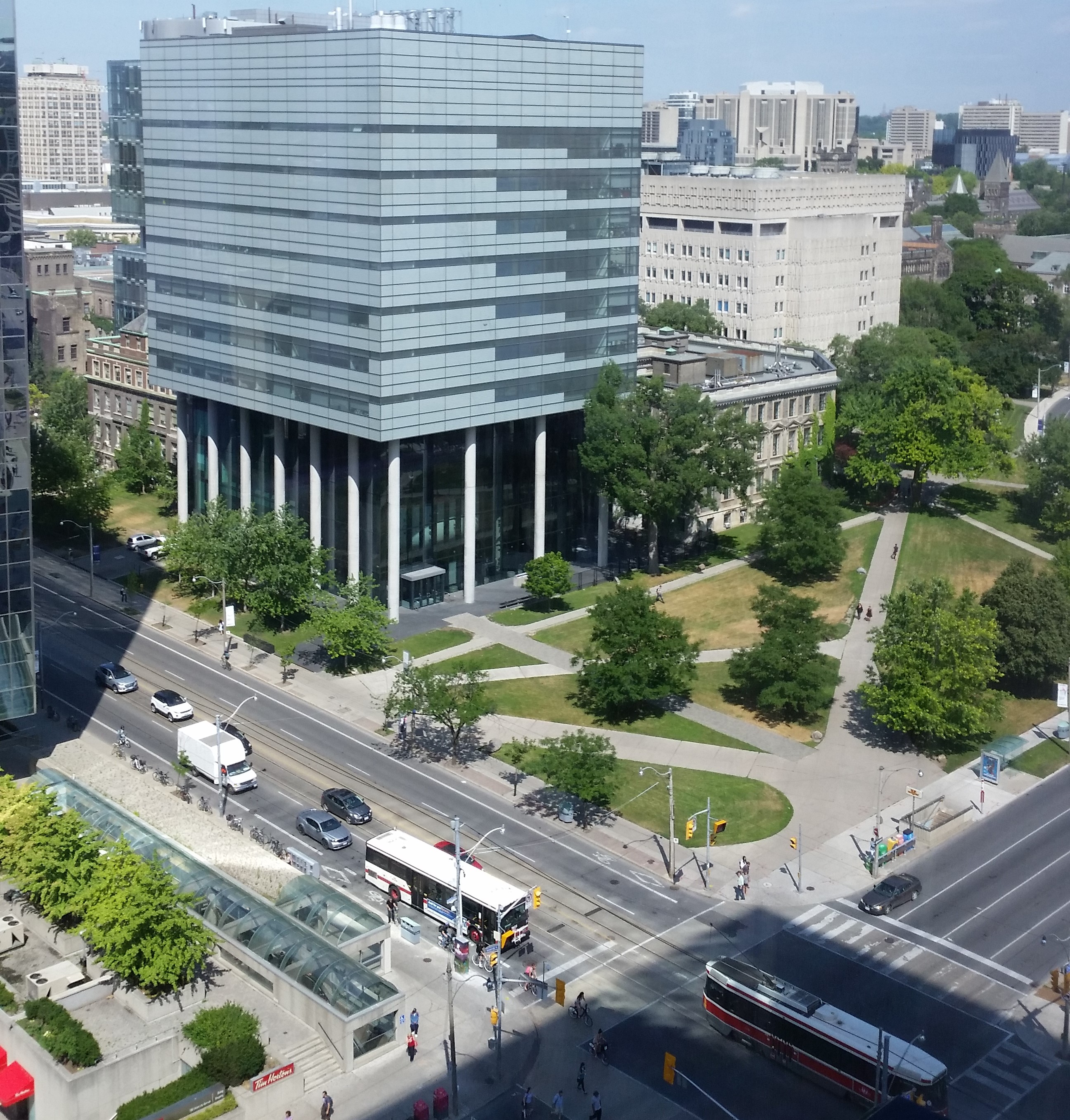
Ryerson Mars: The Botelho Lab is housed in Ryerson MaRS Facilities. These state-of-the art research and study facilities were inaugurated in August 2016 and will bring together about 10 research labs in the Department of Chemistry and Biology. We got the goodies that you need to do your work, places to study and chill, and the views (11th floor) of Queen's Park, downtown and that other University (and the avenue).

Honorary Lab Members




and-TgPH1,fluorescent-biosensors-for-PI(3)P-in-RA- macrophages-pre-treated-with-Dextran(red).png)